Genome Assembly
Sequencing Technologies
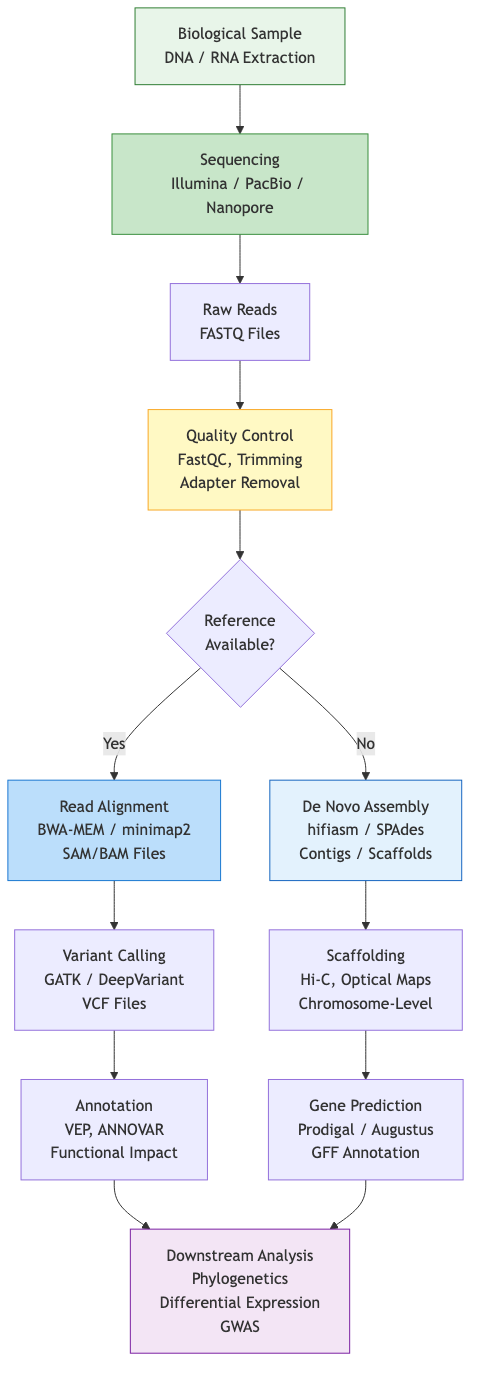
Genome assembly reconstructs contiguous genomic sequences from shorter sequenced fragments. The characteristics of sequencing platforms fundamentally shape assembly algorithms.
Illumina (Short-Read Sequencing)
Sequencing by synthesis on a flow cell. Cluster generation amplifies single molecules via bridge PCR. Fluorescently labeled reversible terminators are incorporated one base at a time, imaged, then cleaved.
- Read length: 75-300 bp (paired-end: 2x150 most common)
- Error rate: ~0.1-1% (substitution-dominated, quality decreases toward 3' end)
- Throughput: Up to 6 Tb per run (NovaSeq 6000)
- Insert sizes: 300-800 bp (standard), 2-20 kb (mate-pair libraries)
- Cost: ~$5-10 per Gb
Pacific Biosciences (PacBio)
Single-molecule real-time (SMRT) sequencing in zero-mode waveguides. Polymerase incorporates fluorescent nucleotides observed in real time.
- CLR (Continuous Long Reads): 10-100 kb, ~10-15% error rate (random insertion/deletion errors)
- HiFi (Circular Consensus Sequencing): Multiple passes around a circularized template; 10-25 kb reads with >99.9% accuracy (Q30+)
- Throughput: 15-30 Gb per SMRT cell (Revio)
Oxford Nanopore Technologies (ONT)
Measures ionic current changes as single-stranded DNA translocates through a protein nanopore. Base calling via recurrent neural networks (Bonito, Dorado).
- Read length: Routinely 10-100 kb; ultra-long reads >1 Mb possible
- Error rate: ~3-5% (systematic homopolymer errors; improving with R10.4 chemistry and duplex sequencing)
- Throughput: 50-100 Gb (PromethION flow cell)
- Unique features: Direct RNA sequencing, real-time base modification detection, adaptive sampling
Other Technologies
- Hi-C: Chromatin conformation capture; provides long-range (Mb-scale) proximity information for scaffolding and phasing
- Linked reads (10x Genomics, discontinued): Short reads with barcode tags from same long molecule
- Optical mapping (Bionano): Restriction enzyme site patterns on long DNA molecules
Assembly Graph Theory
Overlap-Layout-Consensus (OLC)
Classical approach suited for long reads (Sanger, PacBio CLR, Nanopore):
- Overlap: Find all pairwise overlaps between reads using all-vs-all comparison (minimap2, DALIGNER). Store as overlap graph where nodes = reads, edges = overlaps.
- Layout: Identify a consistent path through the overlap graph (Hamiltonian path problem). Remove transitive edges, resolve bubbles from heterozygosity and errors.
- Consensus: Generate consensus sequence from the layout using multiple alignment or statistical models.
Complexity: The overlap step is O(n^2) in reads; heuristic filtering (minimizers, seed-and-chain) makes this tractable. The layout phase must handle repeat-induced ambiguities.
De Bruijn Graph Assembly
Dominant approach for short-read assembly:
- Decompose reads into all k-mers (subsequences of length k)
- Construct a de Bruijn graph: nodes are (k-1)-mers, edges represent k-mers connecting consecutive (k-1)-mers
- Find an Eulerian path through the graph (polynomial time, unlike the Hamiltonian path in OLC)
Key parameters and challenges:
- k-mer size: Small k increases connectivity but introduces ambiguity at repeats; large k resolves repeats but fragments the graph at low-coverage regions. Typical range: 21-127.
- Multi-k assembly: SPAdes uses multiple k values and merges resulting graphs
- Error handling: Erroneous k-mers create tips (dead-end paths) and bubbles; removed by coverage-based filtering
- Repeat resolution: Repeats longer than k create tangles; paired-end information and coverage depth help resolve them
String Graph
A variant of OLC where reads are nodes but transitive overlaps are removed, yielding a compact graph. Used by assemblers like SGA and hifiasm. Conceptually cleaner than de Bruijn graphs for long reads since it avoids k-mer decomposition artifacts.
Assemblers
SPAdes (Short Reads)
Multi-k de Bruijn graph assembler with error correction:
- BayesHammer: Read error correction using Bayesian subclustering of k-mer neighborhoods
- Graph construction: Build de Bruijn graphs for multiple k values; merge into a unified assembly graph
- Paired-end resolution: Use paired reads and coverage to resolve repeats
- Mismatch and chimera correction: Post-assembly polishing
Variants: metaSPAdes (metagenomes), rnaSPAdes (transcriptomes), coronaSPAdes (viral genomes).
Hifiasm (HiFi Long Reads)
Produces phased assemblies from PacBio HiFi reads:
- All-vs-all overlap using minimizer-based seeding
- Haplotype-aware string graph construction: preserves both haplotypes as separate paths
- Graph cleaning: Removes errors while retaining genuine heterozygosity
- Trio binning (optional): Uses parental short reads to phase haplotypes
Produces near-complete, highly accurate diploid assemblies. Telomere-to-telomere assemblies are now routine for human-sized genomes with sufficient HiFi + ultra-long ONT coverage.
Other Notable Assemblers
- Canu: OLC assembler for noisy long reads; correction + trimming + assembly pipeline
- Flye: Repeat graph assembly for long reads; handles high-error data well
- MEGAHIT: Memory-efficient de Bruijn graph assembler for metagenomes using succinct data structures
- Verkko: Hybrid assembler combining HiFi and ultra-long ONT reads for telomere-to-telomere assembly
- wtdbg2: Fuzzy de Bruijn graph approach for ultra-fast long-read assembly
Scaffolding and Gap Filling
Scaffolding
Scaffolding orders and orients contigs into larger scaffolds using long-range information:
- Mate-pair/linked reads: Bridge gaps of known approximate size
- Hi-C: Chromatin proximity ligation data; SALSA2, 3D-DNA, YaHS use contact frequency (decaying as ~1/distance) to order contigs into chromosome-scale scaffolds
- Optical maps: Provide restriction site pattern scaffolding (Bionano Solve)
- Genetic/physical maps: Independent positional information
Gap Filling
Fills N-runs in scaffolds using reads spanning the gap (GapFiller, TGS-GapCloser for long reads).
Polishing
Correct residual errors in consensus sequences:
- Short-read polishing: Pilon maps Illumina reads back to assembly; corrects SNPs, small indels
- Long-read polishing: Medaka (ONT), DeepConsensus (PacBio)
- Iterative polishing: Multiple rounds may be needed, especially for ONT assemblies
Assembly Quality Assessment
Contiguity Metrics
- N50: The contig/scaffold length such that contigs of this length or longer cover 50% of the total assembly. Higher is better. NG50 normalizes to expected genome size.
- L50: The number of contigs needed to reach 50% of total assembly length
- Largest contig: Single longest assembled sequence
- Total span: Sum of all contig lengths; should approximate expected genome size
- Gap content: Percentage of N bases in scaffolds
Completeness Metrics
BUSCO (Benchmarking Universal Single-Copy Orthologs): Searches the assembly for conserved single-copy genes expected in the taxonomic lineage. Reports percentages as Complete (single + duplicated), Fragmented, and Missing. A high-quality assembly typically shows >95% complete BUSCOs.
k-mer completeness (Merqury): Compares k-mer spectrum of the assembly to that of raw reads. Measures QV (consensus quality value), completeness, and phasing accuracy without requiring a reference genome.
Correctness Assessment
- QUAST: Compares assembly to a reference genome; reports misassemblies, mismatches, indels, NGA50
- Inspector: Long-read based assembly evaluation detecting structural and base errors
- Read mapping rate: Fraction of reads that align back to the assembly
Metagenome Assembly
Metagenome assembly reconstructs genomes from mixed microbial communities, introducing unique challenges:
- Uneven coverage: Dominant species at 1000x, rare species at 0.1x
- Inter-species repeats: Conserved genes create chimeric assemblies between related organisms
- Strain variation: Closely related strains create tangles in the assembly graph
Approaches
- Co-assembly: Pool samples for better coverage; risk of chimeras increases
- Sample-specific assembly: Reduces chimeras but may fragment low-abundance genomes
- Binning: Group contigs into metagenome-assembled genomes (MAGs) using composition (tetranucleotide frequency, GC content) and differential coverage across samples (MetaBAT2, MaxBin2, CONCOCT). DAS Tool integrates multiple binning results.
MAG Quality Standards (MIMAG)
- High quality: >90% completeness, <5% contamination, presence of 23S/16S/5S rRNA and >18 tRNAs
- Medium quality: >50% completeness, <10% contamination
- Assessed by CheckM/CheckM2 using lineage-specific marker gene sets
Current State and Challenges
The T2T (Telomere-to-Telomere) Consortium completed the first gap-free human genome in 2022, resolving centromeres, segmental duplications, and rDNA arrays. Key remaining challenges include polyploid genome assembly (e.g., hexaploid wheat), highly repetitive genomes, population-scale pangenome construction (via minigraph-cactus), and real-time assembly for clinical and field applications.